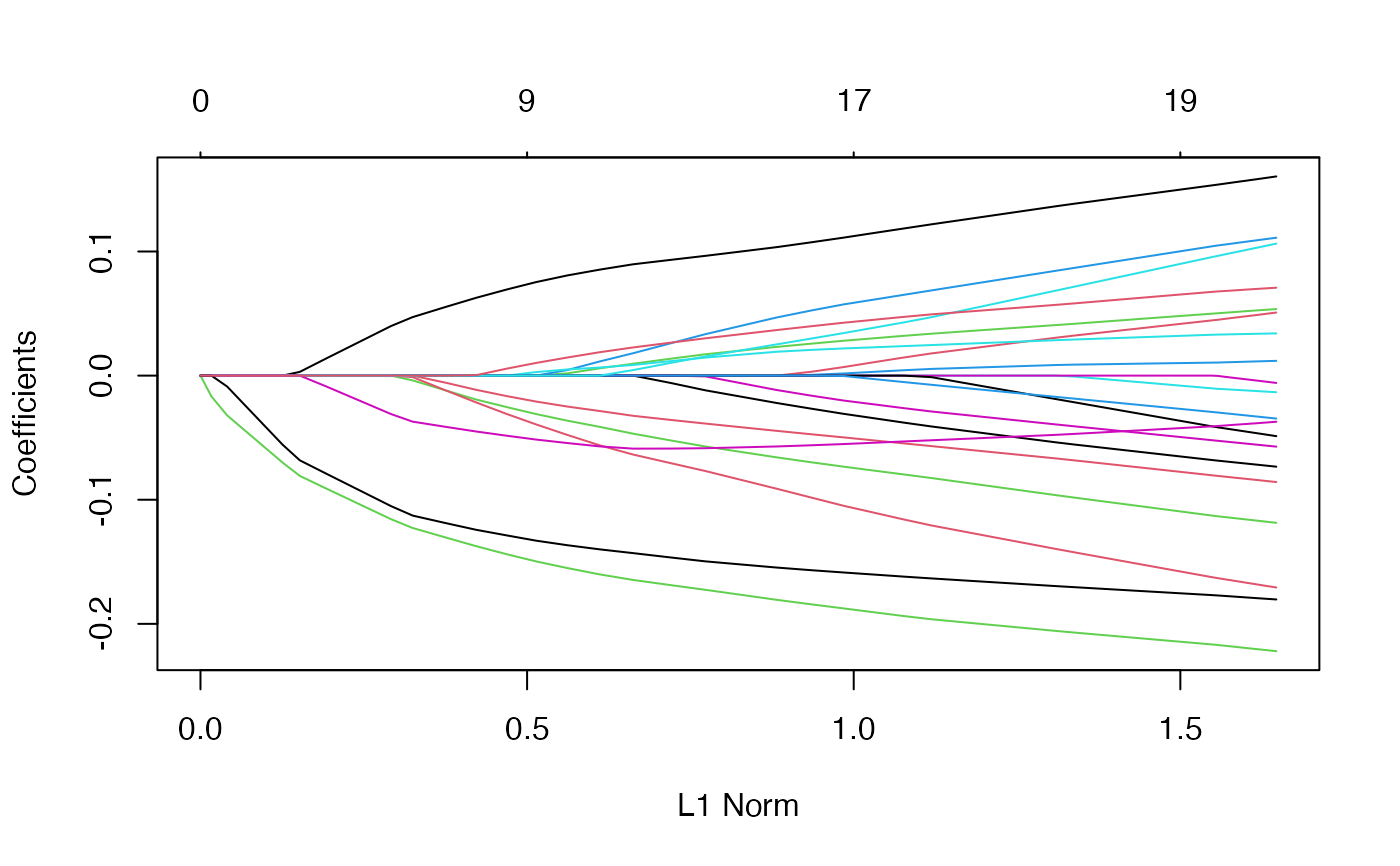
plot coefficients from a "glmnet" object
plot.glmnet.RdProduces a coefficient profile plot of the coefficient paths for a fitted
"glmnet" object.
# S3 method for glmnet
plot(x, xvar = c("norm", "lambda", "dev"), label = FALSE, ...)
# S3 method for mrelnet
plot(
x,
xvar = c("norm", "lambda", "dev"),
label = FALSE,
type.coef = c("coef", "2norm"),
...
)
# S3 method for multnet
plot(
x,
xvar = c("norm", "lambda", "dev"),
label = FALSE,
type.coef = c("coef", "2norm"),
...
)
# S3 method for relaxed
plot(x, xvar = c("lambda", "dev"), label = FALSE, gamma = 1, ...)Arguments
- x
fitted
"glmnet"model- xvar
What is on the X-axis.
"norm"plots against the L1-norm of the coefficients,"lambda"against the log-lambda sequence, and"dev"against the percent deviance explained.- label
If
TRUE, label the curves with variable sequence numbers.- ...
Other graphical parameters to plot
- type.coef
If
type.coef="2norm"then a single curve per variable, else iftype.coef="coef", a coefficient plot per response- gamma
Value of the mixing parameter for a "relaxed" fit
Details
A coefficient profile plot is produced. If x is a multinomial model,
a coefficient plot is produced for each class.
References
Friedman, J., Hastie, T. and Tibshirani, R. (2008) Regularization Paths for Generalized Linear Models via Coordinate Descent
See also
glmnet, and print, predict and coef
methods.